Code Change
By Allison Fromme
A single sick 12-year-old quarter horse mare caught Julia Felippe’s attention in 2001.
As a Ph.D. student at the College of Veterinary Medicine, she reviewed the case of the horse suffering from recurrent pneumonia and noticed something strange: the aggressive proliferation of a common but usually harmless bacterium. Felippe realized that the horse’s immune system must be compromised. Further tests revealed that an important immune component – B-cells – were missing. And yet, other immune cells were present and accounted for.
The fatal disease, which was remarkably similar to a human version, had never been described in horses before. Felippe named it Common Variable Immunodeficiency (CVID) and published the index case, thinking she’d never see another. But in the years since, and now as a faculty member in the Department of Clinical Sciences, she’s uncovered 54 additional cases and embarked on a quest to understand the molecular mechanisms underlying the disease.

“The only way we could study this disease was to work with field veterinarians to understand how it manifested clinically and then investigate the basic mechanisms of the disease in the lab,” she says.
What could cause such a specific deficiency? A virus? Or something in the environment? But no, these searches turned up nothing. Perhaps a genetic mutation, altering the DNA? No, not that either.
Perhaps, then, the culprit is a devastating modification to the way DNA is interpreted by the cell. A change that doesn’t mutate the DNA sequence itself but does alter how cellular machinery interacts with it. An epigenetic cause.
Beyond DNA
Epigenetics is an area of intense research at the college that promises to illuminate basic biology and revolutionize human and animal medicine. Felippe and her colleagues – including Drs. Scott Coonrod, Charles Danko, Praveen Sethupathy ’03 and Paul Soloway ’79 – are making key cell biology discoveries across diverse diseases and conditions, identifying disease biomarkers, characterizing new drug targets – and innovating new methods to make such breakthroughs.
“DNA influences many of the traits we have throughout our lives,” says Soloway, chair of the Department of Biomedical Studies. “But that’s not all. There’s another level of information that we carry. Chemical modification to the DNA, and the proteins that give our chromosomes structure, also influence our traits. And those chemical modifications can come and go. They can respond to environmental changes.”
Epigenetics, in its broadest sense, encompasses the factors beyond DNA that influence which genes are turned off or on, dialed up or dialed down, in a process called transcription. What does that look like? The addition of a carbon atom, or methyl group, on DNA might interfere with the binding of a transcription factor required to express a gene. Structures known as histones can tightly coil segments of DNA, effectively hiding genes. Strands of RNA, DNA’s cousin, can bind to DNA and activate genes. Epigenetics explains how all our cells – from brain cells to blood cells – can contain the same DNA, and yet look and act so differently.
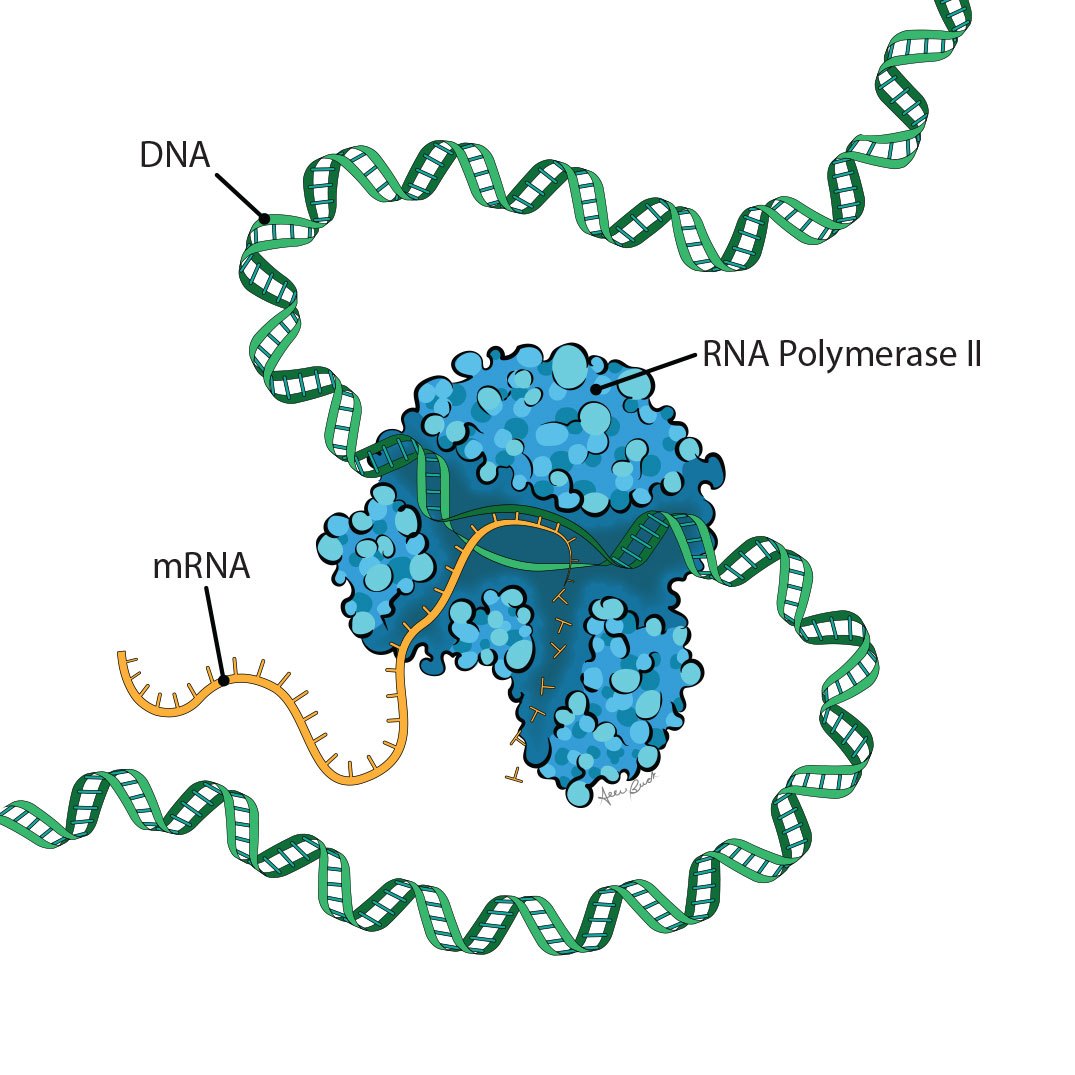
“When you start to look at all of those different modifications, you begin to see how incredibly complex gene regulation is,” says Coonrod, the Judy Wilpon Professor of Cancer Biology at the Baker Institute for Animal Health.
Plus, molecular factors like these can change in response to external inputs like infections, injuries, toxins and nutrients. Understanding these factors – and attempting to manipulate them for health benefits – forms the basis of this field.
Felippe began to investigate different epigenetic mechanisms that could help explain CVID. One analysis of CVID horse samples revealed where methyl groups were attached to DNA throughout the entire genome.
“Most of the epigenetic aberrations we observed were related to B-cells and the expression of antibodies or immunoglobulin,” she says. “So this told us that whatever triggers common immunodeficiency to start with, one of the effects is shutting down the transcription of genes using methylation or hypermethylation. And the more we know about the mechanism, the better we can target it.”
Felippe would ultimately like to develop a therapy for the disease, and she has attempted to reverse methylation using a drug treatment. But it’s still a work in progress.
“This disease teaches us about what we call hematopoiesis, how cells are produced in the bone marrow. What are the steps, the mechanisms, that need to be in place? It teaches us specifically about hematopoietic stem cells, and that’s important because we are in a field that is very thirsty, waiting for discoveries,” she says. “And our ability to edit epigenetic conditions is going to be a common thing in the future.”
Unexplored elements
The human brain represents some of the most unexplored elements in biology, says Soloway. In a collaboration with Barbara Strupp, Ph.D. ’82, professor of nutrition in the College of Human Ecology, he’s exploring the possibility that epigenetic mechanisms might explain cognitive differences characteristic of Down syndrome. Strupp previously demonstrated that the essential nutrient choline positively influences behavior in a mouse model of Down syndrome.
“Yes, we can identify differences. The next question is what do those differences mean? Can they help explain behavioral differences?”
Dr. Paul Soloway
Might mice with Down syndrome have a different epigenetic profile compared with unaffected mice? To answer this question, Soloway’s graduate student, Roman Spektor, used a technique called single cell ATAC-seq to analyze thousands of individual brain cells from mice with and without Down syndrome.
“This method is really powerful,” says Soloway, because lots of different cell types make up brain tissue. If the tissue is analyzed whole, signals that are highly relevant in one cell type are lost when averaged with other types.
Across different brain cells, Soloway mapped the locations of histones – the structures that wrap up DNA, interact with each other and form the basis of DNA’s three-dimensional chromatin structure. And that structure dictates which genes are available for transcription.
“Yes, we can identify differences. The next question is what do those differences mean? Can they help explain behavioral differences?” Soloway asks.
Next, Soloway plans to compare Down syndrome mice that have had choline supplementation with those that haven’t. He hypothesizes that the nutrient could alter chromatin structure, a finding that would have important health implications.
Chemical changes
Just up the hill at the Baker Institute for Animal Health, Coonrod is studying histones in the context of breast cancer.
“Estrogen signaling is the main driver of most breast cancers. A simple way to think about it is this: estrogen binds to estrogen receptors, which regulate a whole suite of genes, many of which are cell proliferation genes,” Coonrod explains. And when that pathway becomes hyperactivated, tumors grow.

Even when DNA is tightly wrapped around histones, estrogen receptors can bind to it, but only weakly. Because the DNA isn’t fully available, genes aren’t immediately transcribed. Coonrod’s evidence suggests that estrogen receptors recruit a special enzyme, PAD2, to help. PAD2 chemically changes the histone, loosening its grip on the DNA and allowing the expression of genes – the same genes implicated in tumor growth.
Coonrod expects that understanding this mechanism will one day help patients whose breast cancer is resistant to tamoxifen, the drug commonly used to prevent recurrence after surgical tumor removal. He hypothesizes that high PAD2 levels could indicate that the tumor is likely to become tamoxifen-resistant and alert doctors to offer different therapies to such patients. In this way, PAD2 holds promise both as a clinical biomarker and as a target for future drug development.
Addicted to an altered microenvironment
Sethupathy, associate professor in the Department of Biomedical Sciences, is taking a genome-wide approach to studying fibrolamellar carcinoma (FLC), a rare but devastating liver cancer affecting children and young adults.
“There’s a growing sentiment in the cancer field in general that cancers become addicted to an altered microenvironment, and usually what that means is an altered chromatin environment,” Sethupathy says.
“We wanted to identify how the chromatin environment is altered across the entire genome in FLC patients, comparing their own cancerous tissue to neighboring nonmalignant tissue,” he says. “And what we’ve found is that there are hotspots – regions across the genome that are massively flaming signals of hyperactivity. But if you look at the normal liver, those regions are completely inactive.”
One hotspot pointed to a particular lncRNA, a molecule with an unknown role. “We are in the midst of doing functional studies to determine whether shutting down this long noncoding RNA might be sufficient to at least partially mitigate some of the cancer phenotypes, maybe slow its growth, maybe prevent its metastatic capability,” he says.
Valuable techniques
Sethupathy’s study might not have been possible without a Cornell-pioneered technique called ChRO-seq. The liver tissue samples had been frozen, and other methods relied on cells from live cultures.
Danko developed this application of a method initially invented by Hojoong Kwak, Ph.D. ’13, then a graduate student in the lab of Dr. John Lis and now an assistant professor in the Department of Molecular Biology and Genetics.
“We put the tissue in liquid nitrogen and then smash it, and that pulverizes it into a really fine powder,” Danko says. “And then you can use a variety of chemicals to isolate DNA – but not just DNA, DNA and histones, and RNA polymerase itself. Then we can use the same elegant biochemistry that was already cooked up by John Lis’ lab.”
“Knowing what pathways are activated gives a really good source of information for diagnostics.”
Dr. Charles Danko
In a recent study, Danko investigated glioblastoma, an aggressive brain cancer, and demonstrated the value of ChRO-seq. Some of the tissue samples analyzed dated as back as the 1970s and were partially degraded. But RNA polymerase, the machinery that makes RNA, remained attached to specific DNA regions and retained its activity.
“It’s kind of amazing to me, you can actually start it up again and make new RNA,” Danko says. And that process shows which DNA regions are active. “Knowing what pathways are activated gives a really good source of information for diagnostics. With the glioblastoma project, we developed a signature where the transcription factors themselves could predict roughly a two-fold difference between two groups of patients and their outcomes.”
In addition to the potential health applications for his work, Danko’s primary interest is understanding how DNA sequences are used to create different morphologies and biological functions.
“Scientists sequenced the genome decades ago but we still basically have no idea what most of it does and how it works together. From a basic science perspective, I think that is the most fascinating question in epigenetics right now.”
Illustrations by Allie Buck. Photos by Rachel Philipson, Cornell Brand Communications and the Cornell University College of Veterinary Medicine.


